Update: Please refer to the comments below to read about updated versions of the software (https://www.nitrc.org/frs/?group_id=889). Neuroimaging software is frequently advancing so check with man pages or other documentation for options and changes.
A new folder called 'BeauchampMacMRIcron' will be created. Open the folder; you should see the file mricron.app next to a small brain icon. To start the program, double click on the mricron.app file or the small brain icon. The dataset name is listed at the bottom of the window for Mac. The correct brain to view is the 'ch2' file (ch2.nii.gz). Mar 01, 2011 MAC gui version is user friendly but it can convert one data set at a time. Therefore, I wrote an applescript to convert multiple DICOM data sets to NIfTI/Analyze format at one instance using dcm2nii command. Click below to download script. MRIcron is a user-friendly and efficient piece of software which aims to assist you in viewing medical imagery with multiple layering, supporting HDR, NII and VOI formats, thus enabling you to.
Note: While I will try to be verbose in my how-to guides of imaging, I assume any readers have at least basic knowledge of the command line (Terminal) and bash. If you feel lost with some of the bash references and need some additional info, here is a good beginner’s guide to bash.
Usually the first step to process structural MRI data is to convert the DICOM files from the MR scanner into a more useable format. The current “best” format for files is NIfTI. There are other formats but this is a file format that is widely used and very useable. There are problems with it (specifically with how some software deal handle it but that is a topic for a later time).
One of the best tools to convert DICOM files to NIfTI is Chris Rorden’s dcm2nii (see also his profile on the NITRC website for contact info). This is a command line tool (there is also a graphical user interface version) that wil convert just about any DICOM image to an appropriate NIfTI. dcm2nii is part of the MRIcron package – the most recent version of the software can be downloaded from this link from the NITRC website. There are Linux, Macintosh, and Windows versions of the software (as well as source code if you care to build your own version). This software is available to download for free.
After the software is downloaded and extracted, it can be run from wherever its location is. In other words, this is not “installed” software, it is run from whatever location it is saved to (for example, it could be saved in /Applications/mricronmac/ or in /Users/bbunny/neuroimaging/).
See the screenshot for an example of dcm2nii location.
In order to run dcm2nii from Terminal, move into the directory where dcm2nii is (e.g., using the command line type: cd /Applications/mricronmac/ – again, this should be where the actual dcm2nii program is on your computer).
Next, in Terminal type: ./dcm2nii and press return (enter).
This will bring up some text like this:
It gives an example of how to run the program. Also notice my default preferences. I edited the dcm2nii.ini file to have those defaults because I’ve found that they work the best for the work I do. You can also run dcm2nii from anywhere in Terminal by typing: /[path to mricronmac]/dcm2nii where [path to mricronmac is replaced with the full path to where dcm2nii is – e.g., /Applications/mricronmac/ (check out this post to see how to be able to run a program without typing in the whole path to it). I’ve found it easiest to run dcm2nii from within the folder of DICOM files I want to convert to NIfTI images. To do this first you need to move to the directory where your raw DICOMs are (I typically make a temporary copy of the files so I am not doing anything with the originals, just to prevent any potential issues): for example, /Users/Shared/DWI/blindnum001/DATA/64dir_DWI. Now from within this directory you can run dcm2nii and convert the images with the following command (or whatever is the equivalent on your computer):
Micron Macro Trends
/Applications/mricronmac/dcm2nii –a y –d n –e n –f y –g y –i n –r n –x n /Users/Shared/DWI/blindnum001/DATA/64dir_DWI/image001.dcm
Note: you do not need to include all those options (–a y –d n –e n –f y –g y –i n –r n –x n), I included them just to show what are the defaults that I run with dcm2nii. If you have edited the dcm2nii.ini file or if you want to use other defaults or options, go ahead – those are just the options I prefer. After the conversion completes you should have one or more NIfTI files in the same directory where your dicoms are. For diffusion weighted data you should have three files: the .nii.gz (if you selected the gzip option), a .bval, and a .bvec. The only times I’ve not had dcm2nii create bval or bvec files (and I have verified that the dicom files are not corrupted) is when I did not change the default options for dcm2nii, which resulted in overly long NIfTI file names; that, for some reason, prevented the bval and bvec files from being created. That is why I make sure output filename is only the input filename. I’ve never had any conversion issues since setting that option.
There is your conversion. You are all done with this step of neuroimaging!
Alternatively, you can use the graphical user interface for dcm2nii:
First you want to make sure your preferences look something like this (if these are the preferences you want; I prefer just to keep the input filename because it tends to be descriptive but fairly short; I typically rename my files later):
In the GUI under File select DICOM to NifTi and locate the dicoms you wish to convert in the dialogue box that appears. After you select the files and push Open the conversion will run. When it is finished verify that you have a .nii.gz file and .bvec and .bval files. The bval file is a simple text file that includes all the b-values of the diffusion scan (e.g., for a 64 direction scan with one b0 image there will be 65 numbers in there – one 0 and 64 1000s or whatever your b weighting was for your scan. 1000 is pretty typical). I prefer using the command line because it can speed up the process but either way will result in NIfTI images.
Now we are ready to move on to another step in image processing.

Micron Machine Tools
This is the web page for information about downloadable MRI neuroanatomy teaching materials. The instructions below describe how to download MRI datasets and a freely available viewing program to look at them.
REQUIRED Step 1: Download MRIcroN (free software for viewing MRI datasets)Download the correct installer package for your computer. Windows, Mac and Linux are supported; detailed installation instructions are provided for Windows and Mac but not Linux. For Windows:The following instructions were tested with Windows 7 running Internet Explorer 9. For different version of Windows or other web browsers, the steps may be slightly different.
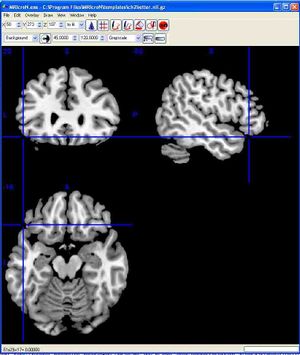
After successfully starting the program, you should see the following window. To make the images easier to see, you can enlarge the window. Go to Step 2, below. Windows MRIcroN window For Macintosh:The following instructions were tested with Mac OSX 10.8.2 running Safari. For different versions of OSX or other web browsers, the steps may be slightly different.
You should see the following window. To make the images easier to see, you can enlarge the window. To see only one plane enlarged, click View/Axial Only. Macintosh MRIcroN window For Linux (or if the above instructions don't work): please go tohttp://www.mccauslandcenter.sc.edu/mricro/mricron/install.htmlYou are on your own for installation! REQUIRED Step 2: Explore the brain and learn structures found in labs 8 and 9Regardless of the platform (PC or Mac), the top left of the window shows a coronal view of the brain. The top right of the window shows a sagittal view of the brain. The bottom left of the window shows an axial (horizontal) view of the brain. Click the mouse at any brain location to move the blue viewing crosshairs.The dataset name is listed at the top of the MRIcroN window in Windows, at the bottom of the window for Mac. The correct brain to view is the 'ch2' file (ch2.nii.gz). Load this file by selecting 'File/Open/Templates/ch2.nii.gz'.At the top of the program window are three boxes labelled 'x', 'y' and 'z' that show the brain location. Enter each of the co-ordinates (x,y,z) listed below into MRIcron. This will take you to the structure whose name is on the first column.This material will make up part of the testing materials for these laboratories (lab exams). Name of structure / Coordinates of structure (enter into x,y,z boxes in MriCron)
Structures in lab 11Limbic System
Teaching Aid: Colored Overlay to show different anatomical regionsWhile the above co-ordinates show examples of single locations and their label, it can also be nice to see the complete extent of an area. This can be done as follows:
MRIcroN with anatomical labels Surf around the brain. When clicking on a colored region, you will see the anatomical label for that colored area of the brain. The label (e.g. Putamen_L for left putamen) will appear in the bottom of the viewing window on PC, in the top of the viewing window on Mac. OPTIONAL Step 3: Different kinds of brain MRIMRI is clinically very useful because it is non-invasive and safe. Because no exposure to radiation is involved, patients can receive many different kinds of MRI imaging procedures without risk. The MRI scanner can be programmed to create different kinds of images that reveal different tissue properties. The three main kinds of images are T1, T2 and Proton Density (FLAIR). The names of the image types refer to the physical priniciples used to create the images. The three links below link to three datasets, all collected from the same normal subject, that show the three different image types. Click on each link and save the file to your computer. After you have downloaded the datasets, you can load and view them in MriCroN by clicking File/Open. Navigate to the directory where you saved them and select the dataset name. OPTIONAL Step 4: Different kinds of spine MRIBecause the spine is so long, it is typically imaged in three separate scans covering the cervical, thoracic, and lumbar+sacral portions of the cord. The following archive file contains T1, T2 and IR datasets for each spine portion. Download the file and unzip the 12 datasets that it contains. It is easiest if they are unzipped to the same directory as the datasets in Step 3. The datasets can then be opened in MRIcroN. OPTIONAL Step 5: Cerebral VasculatureHere are two movies that show the results of a Magnetic Resonance Angiogram (MRA). Click to download the movies to your desktop. Then double-click on the movies to view them. They have been tested with Apple Quicktime, free. software downloadable from www.apple.com. After the movie is loaded, press the left and right arrow to scroll through the frames of the movie. License Information for MRIcron software.Chris Rorden's MRIcron, copyright 2012, all rights reserved. Redistribution and use in binary forms, with or without modification, are permitted provided inclusion of the copyright notice, this list of conditions and the following disclaimer is provided with the distribution: Neither the name of the copyright owner nor the name of this project (MRIcron) may be used to endorse or promote products derived from this software without specific prior written permission.This software is provided by the copyright holder 'as is' and any express or implied warranties, including, but not limited to, the implied warranties of merchantability and fitness for a particular purpose are disclaimed. In no event shall the copyright owner be liable for any direct, indirect, incidental, special, exemplary, or consequential damages (including, but not limited to, procurement of substitute goods or services; loss of use, data, or profits; or business interruption) however caused and on any theory of liability, whether in contract, strict liability, or tort (including negligence or otherwise) arising in any way out of the use of this software, even if advised of the possibility of such damage. |
Mricron Manual
Retrieved from 'https://openwetware.org/mediawiki/index.php?title=Beauchamp:MedNSLab&oldid=757357'
